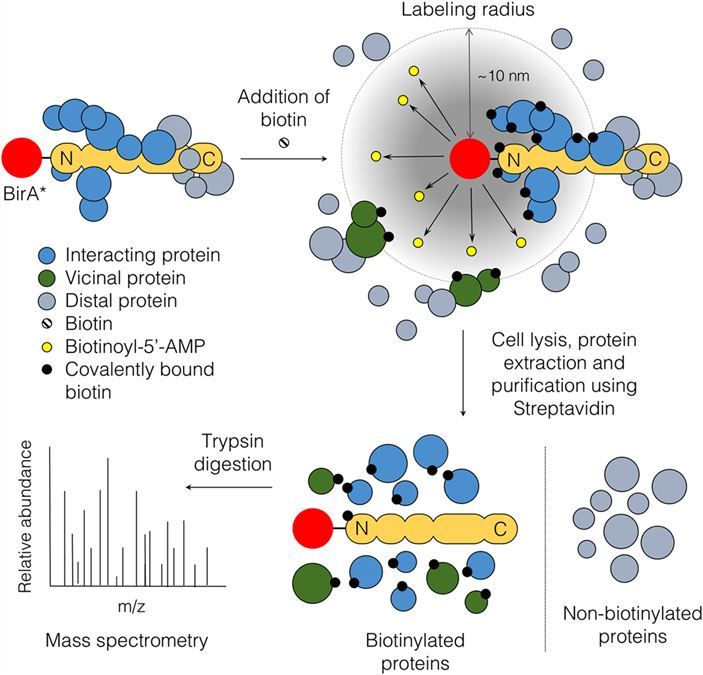
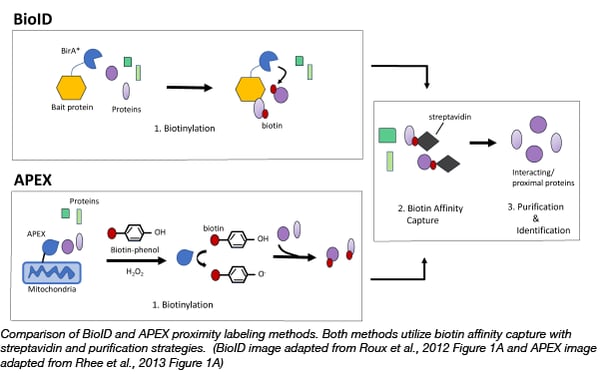
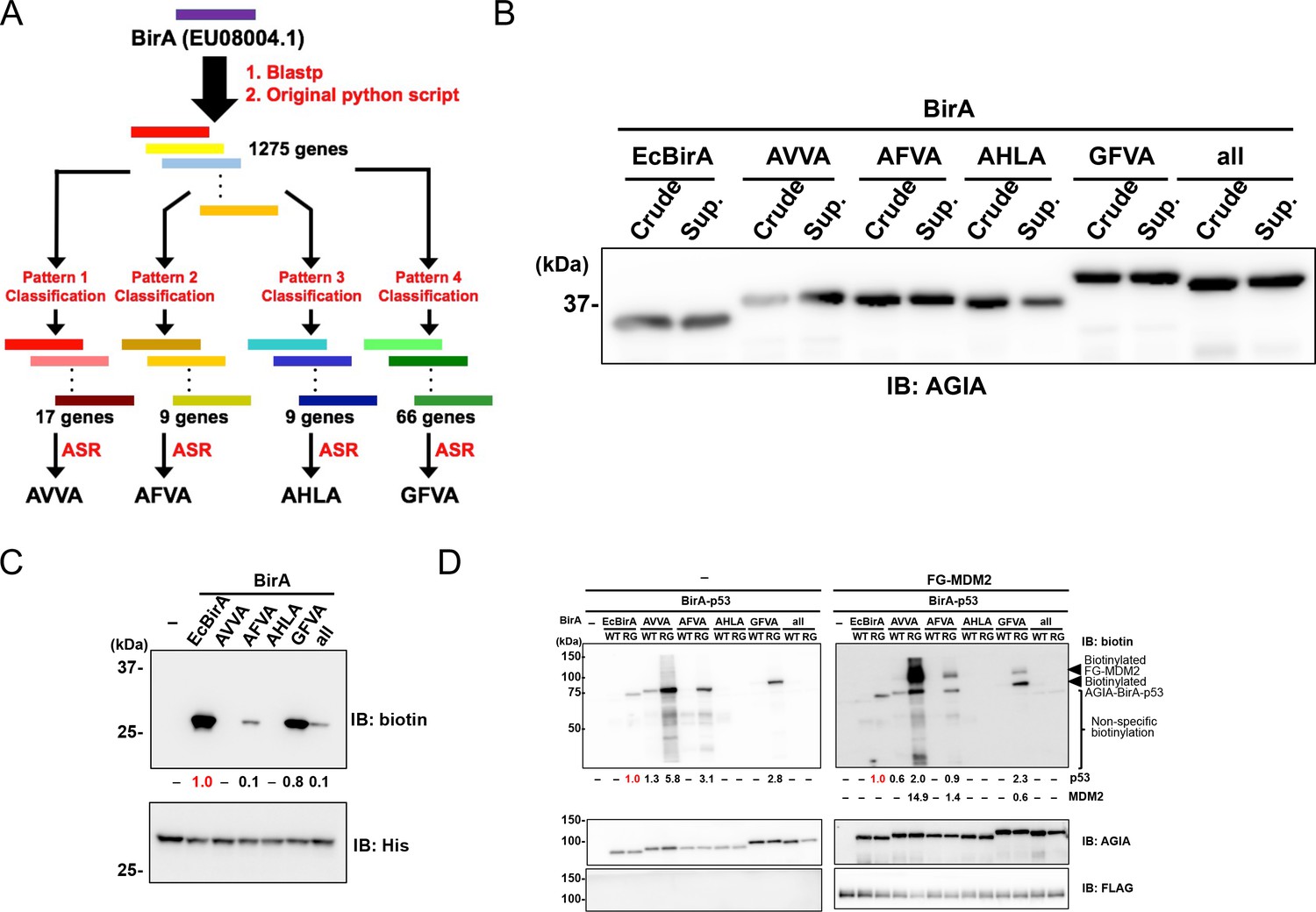
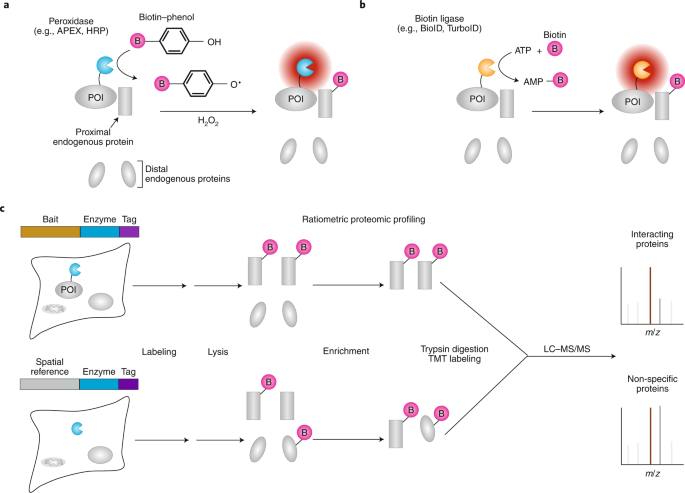
Identification of interacting proteins using proximity-dependent biotinylation with BioID2 in Trypanosoma brucei

A Toolbox for Efficient Proximity-Dependent Biotinylation in Zebrafish Embryos - Molecular & Cellular Proteomics

Mapping protein networks in yeast mitochondria using proximity-dependent biotin identification coupled to proteomics: STAR Protocols

In planta proximity-dependent biotin identification (BioID) identifies a TMV replication co-chaperone NbSGT1 in the vicinity of 126 kDa replicase - ScienceDirect

Identifying Protein-Protein Associations at the Nuclear Envelope with BioID. - Abstract - Europe PMC
![PDF] Probing mammalian centrosome structure using BioID proximity-dependent biotinylation. | Semantic Scholar PDF] Probing mammalian centrosome structure using BioID proximity-dependent biotinylation. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/72e2710524f8b1a033ff55abc8494c4f1e1ccf5f/4-Figure1-1.png)
PDF] Probing mammalian centrosome structure using BioID proximity-dependent biotinylation. | Semantic Scholar
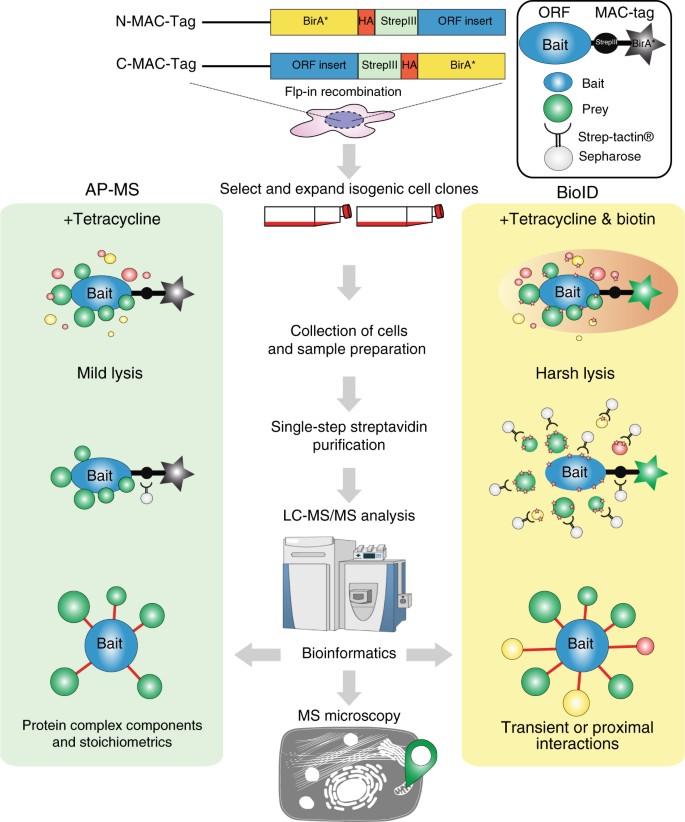
An AP-MS- and BioID-compatible MAC-tag enables comprehensive mapping of protein interactions and subcellular localizations | Nature Communications
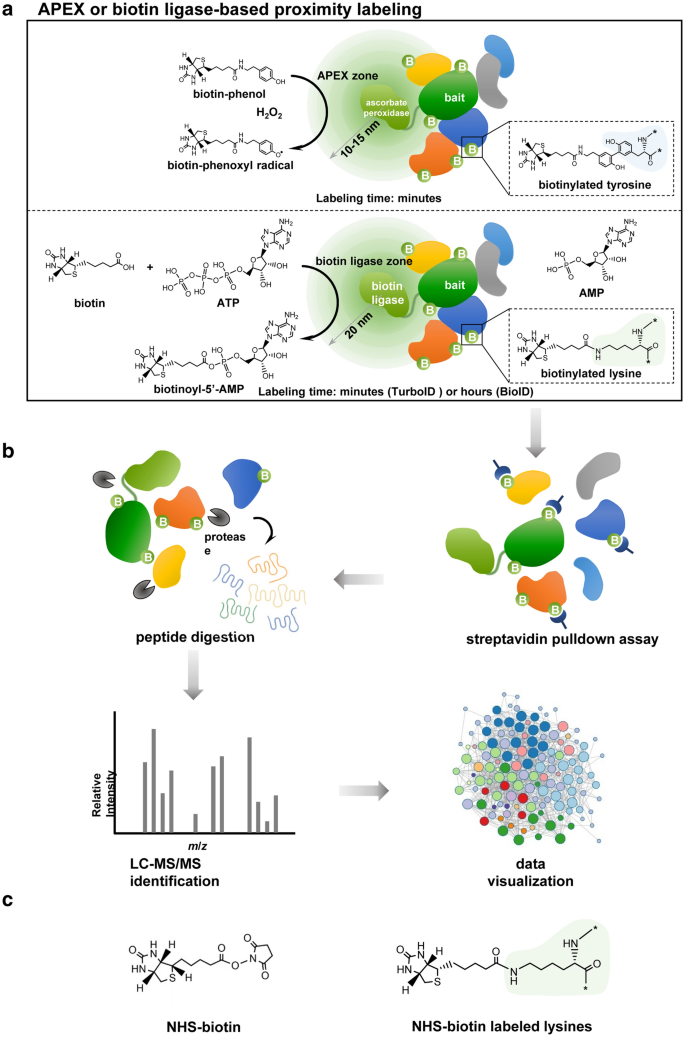
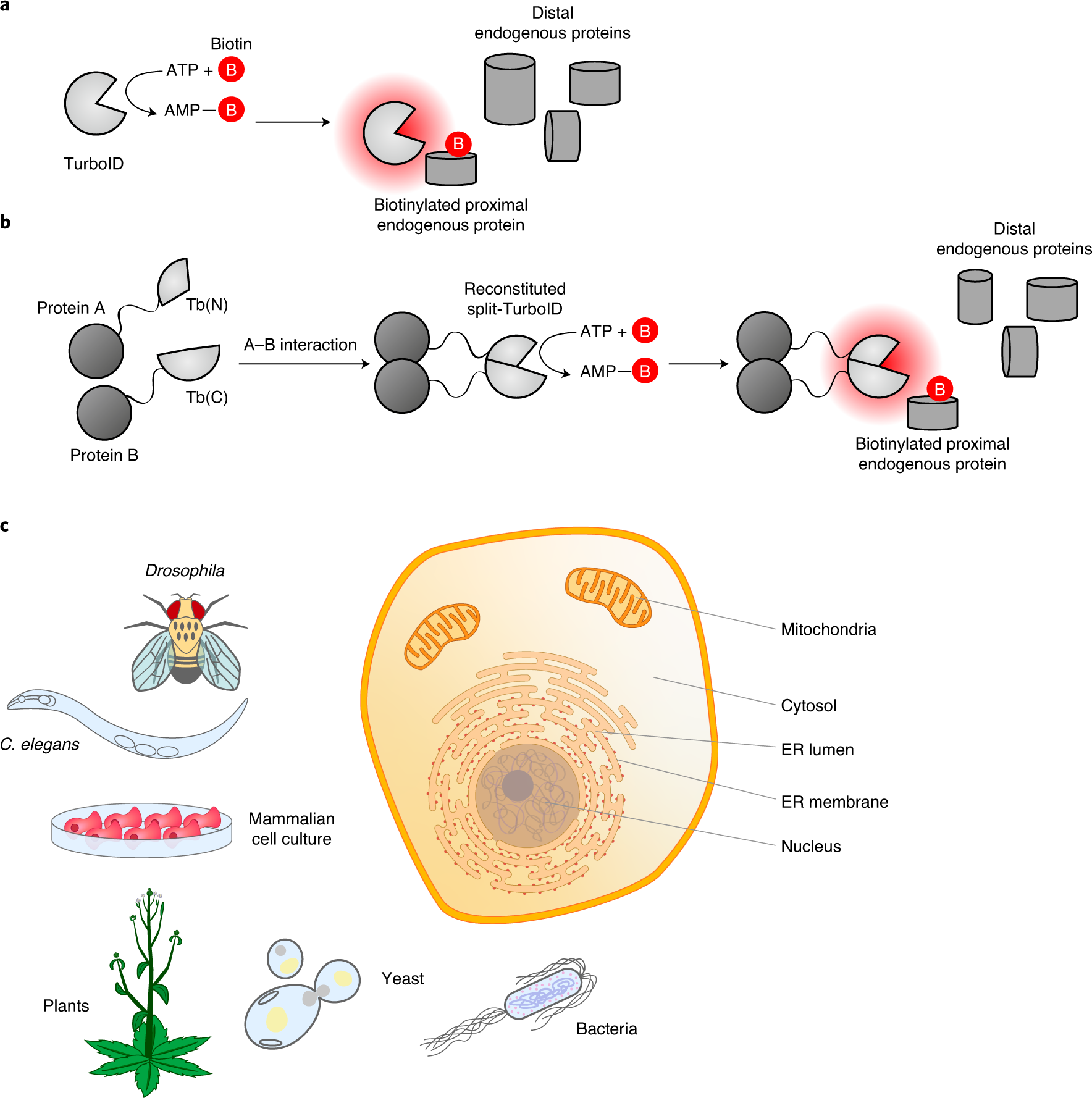
In vivo interactome profiling by enzyme‐catalyzed proximity labeling | Cell & Bioscience | Full Text

BioID Combined with Mass Spectrometry to Study Herpesvirus Protein–Protein Interaction Networks | SpringerLink
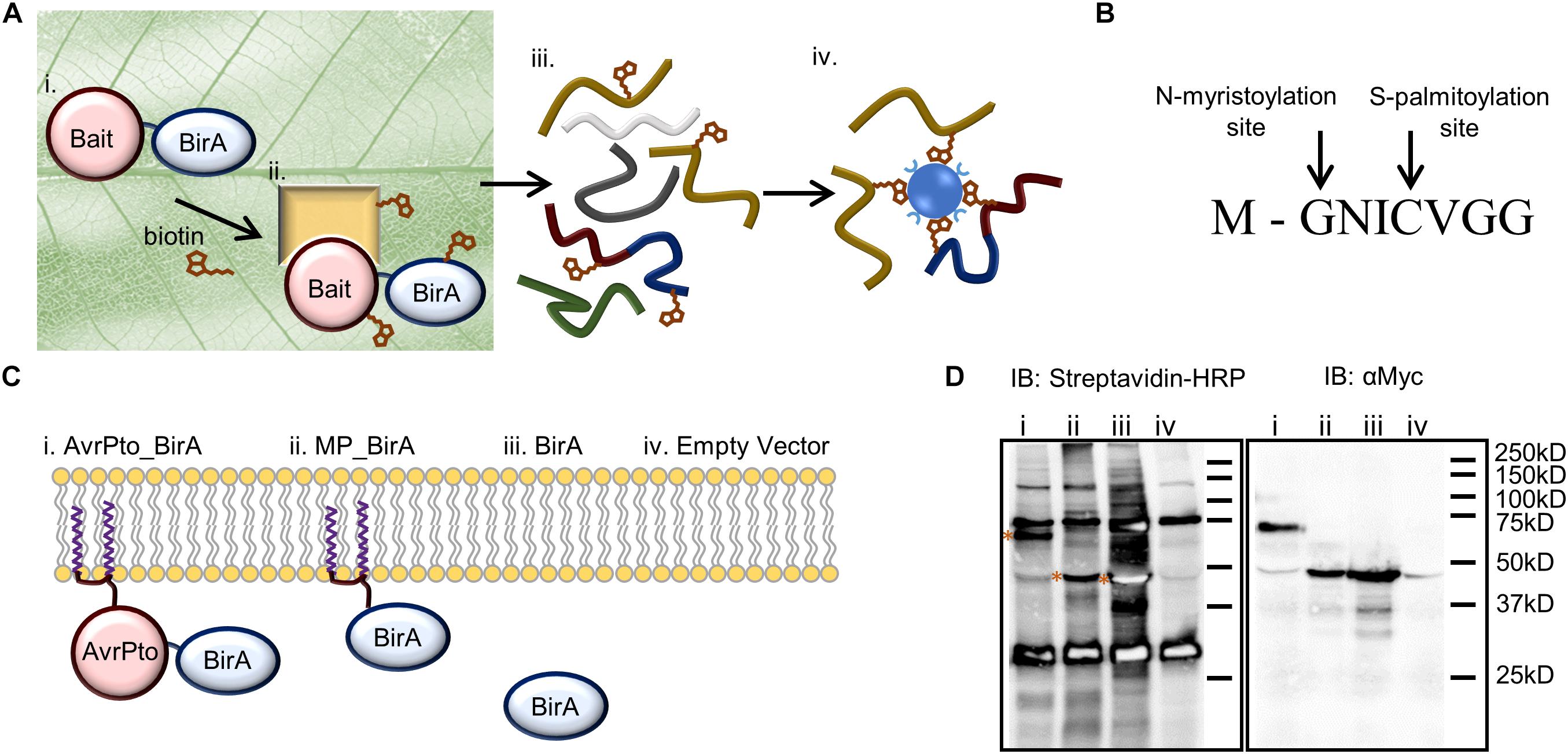
Frontiers | Development of a Rapid in planta BioID System as a Probe for Plasma Membrane-Associated Immunity Proteins